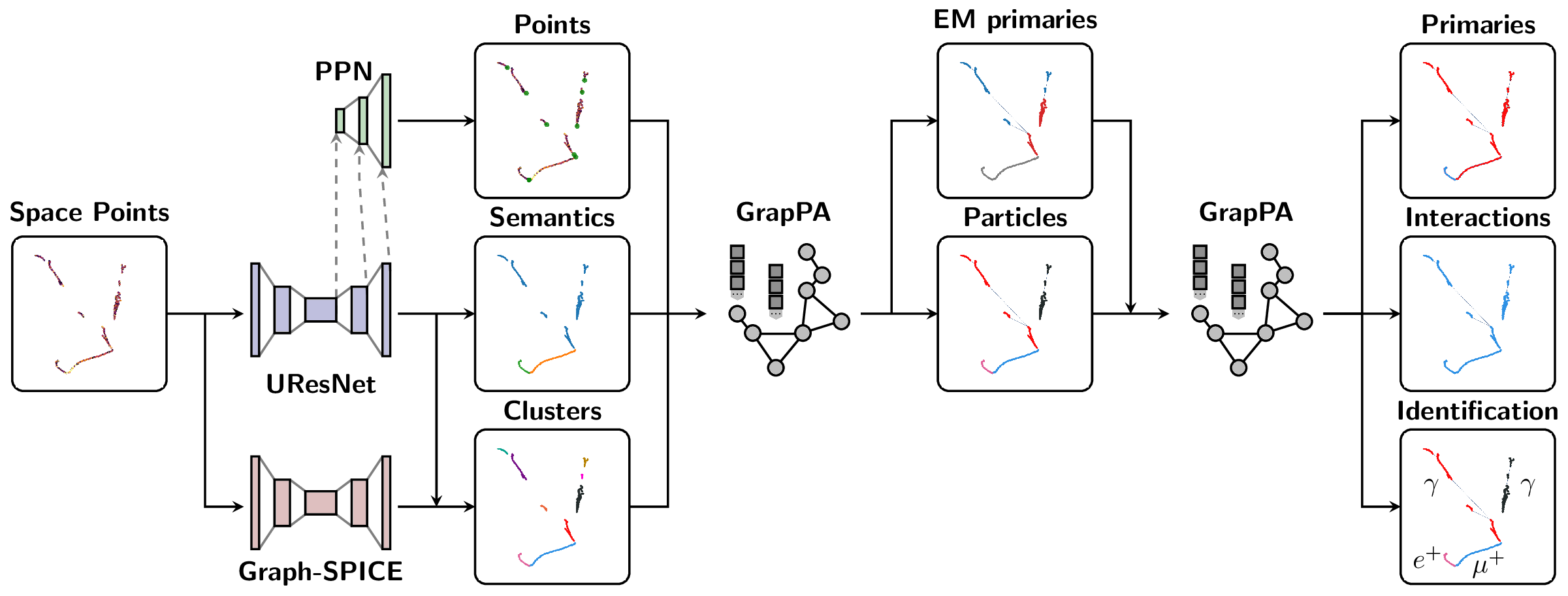
The Scalable Particle Imaging with Neural Embeddings (SPINE) package leverages state-of-the-art Machine Learning (ML) algorithms -- in particular Deep Neural Networks (DNNs) -- to reconstruct particle imaging detector data. This package was primarily developed for Liquid Argon Time-Projection Chamber (LArTPC) data and relies on Convolutional Neural Networks (CNNs) for pixel-level feature extraction and Graph Neural Networks (GNNs) for superstructure formation. The schematic below breaks down the full end-to-end reconstruction flow.
For full SPINE workflows, the recommended runtime is the published SPINE container image released alongside each SPINE version. Use the release-tagged image ghcr.io/deeplearnphysics/spine:<release> when reproducibility matters. When in doubt, use ghcr.io/deeplearnphysics/spine:latest or omit the tag entirely, which is equivalent in Docker-style image references. Docker is the most direct path on workstations and servers; Apptainer/Singularity is the preferred path on HPC systems that do not allow Docker. A local pip installation is mainly intended for post-processing, analysis, visualization, docs, or lightweight development.
SPINE supports both container-based and local Python installation workflows, but they are not equivalent.
Every SPINE release publishes a matching container image to GHCR. For end-to-end reconstruction, training, and inference, use a release tag when you want a pinned environment. When in doubt, use latest or omit the tag entirely:
# Equivalent to: docker pull ghcr.io/deeplearnphysics/spine
docker pull ghcr.io/deeplearnphysics/spine:latest
# Example: replace <release> with a SPINE release tag such as 1.2.3
docker pull ghcr.io/deeplearnphysics/spine:<release>Omitting the tag is equivalent to using latest in Docker-style image references.
Use Docker when you have a local workstation or server with container runtime support:
docker run --gpus all -v $(pwd):/workspace \
ghcr.io/deeplearnphysics/spine:latest \
spine --config /workspace/config/train_uresnet.yaml --source /workspace/data.h5
# Or pin to a specific release
docker run --gpus all -v $(pwd):/workspace \
ghcr.io/deeplearnphysics/spine:<release> \
spine --config /workspace/config/train_uresnet.yaml --source /workspace/data.h5On Apple Silicon macOS systems, the published SPINE image should still be run
as linux/amd64. Specify that explicitly by adding
--platform=linux/amd64 to docker run, for example:
docker run --platform=linux/amd64 --gpus all -v $(pwd):/workspace \
ghcr.io/deeplearnphysics/spine:<release> \
spine --config /workspace/config/train_uresnet.yaml --source /workspace/data.h5For Jupyter notebook/lab use specifically, avoid the Docker Desktop combination of Apple Virtualization Framework with Rosetta enabled: the kernel handshake may stall in that setup even though normal SPINE CLI commands still run. Apple Virtualization Framework without Rosetta and Docker VMM have both been verified to work for Jupyter with the published image.
Use Apptainer or Singularity on HPC systems that do not allow Docker directly. The recommended path is to pull the same released SPINE image from GHCR:
apptainer pull spine_latest.sif docker://ghcr.io/deeplearnphysics/spine:latest
apptainer exec --nv spine_latest.sif \
spine --config /workspace/config/train_uresnet.yaml --source /workspace/data.h5
# Or pin to a specific release
apptainer pull spine_<release>.sif docker://ghcr.io/deeplearnphysics/spine:<release>
apptainer exec --nv spine_<release>.sif \
spine --config /workspace/config/train_uresnet.yaml --source /workspace/data.h5The Docker and Apptainer paths consume the same released image; the difference is only the container runtime.
Use a local pip installation when you only need downstream tooling such as post-processing, analysis, visualization, documentation, or light development.
1. Core Package (minimal dependencies)
# Essential dependencies: numpy, scipy, pandas, PyYAML, h5py, numba
pip install spine2. With Visualization Tools
# Adds plotly, matplotlib, seaborn for data visualization
pip install spine[viz]3. Development Environment
# Adds testing, formatting, and documentation tools
pip install spine[dev]4. Everything (except PyTorch)
# All optional dependencies (visualization + development tools)
pip install spine[all]The published SPINE image already includes the compatible PyTorch, torch-geometric, MinkowskiEngine, and LArCV stack. Use the release-tagged image through Docker or Apptainer as shown above.
# Step 1: Install PyTorch with CUDA
pip install torch --index-url https://download.pytorch.org/whl/cu118
# Step 2: Install ecosystem packages (critical order)
pip install --no-build-isolation torch-scatter torch-cluster torch-geometric MinkowskiEngine
# Step 3: Install SPINE
pip install spine[all]Why the container is preferred: the PyTorch ecosystem (torch, torch-geometric, torch-scatter, torch-cluster, MinkowskiEngine) forms an interdependent group requiring exact version compatibility and complex compilation. The released SPINE container pins that stack for you.
LArCV2 is already bundled in the published SPINE image.
# Clone and build the latest LArCV2
git clone https://github.com/DeepLearnPhysics/larcv2.git
cd larcv2
# Follow build instructions in the repositoryNote: Avoid conda-forge larcv packages as they may be outdated. Use the released SPINE container or build LArCV2 from the official source.
For developers who want to work with the source code:
git clone https://github.com/DeepLearnPhysics/spine.git
cd spine
pip install -e .[dev]For rapid development and testing without reinstalling the package:
# Clone the repository
git clone https://github.com/DeepLearnPhysics/spine.git
cd spine
# Install only the dependencies (not the package itself)
# Or alternatively simple run the commands inside the above container
pip install numpy scipy pandas pyyaml h5py numba psutil
# Run directly from source
python src/spine/bin/run.py --config config/train_uresnet.yaml --source /path/to/data.h5
# Or make it executable and run directly
chmod +x src/spine/bin/run.py
./src/spine/bin/run.py --config your_config.yaml --source data.h5💡 Development Tip: This approach lets you test code changes immediately without reinstalling. Perfect for rapid iteration during development.
To build and test packages locally:
# Build the package
./build_packages.sh
# Install locally built package
pip install dist/spine-*.whl[all]Option 1: Run from the released container:
docker run --gpus all -v $(pwd):/workspace \
ghcr.io/deeplearnphysics/spine:<release> \
spine --config /workspace/config/train_uresnet.yaml --source /workspace/data.h5Option 2: After installation, use the spine command locally:
# Run training/inference/analysis
spine --config config/train_uresnet.yaml --source /path/to/data.h5Option 3: Run directly from source (development):
# From the spine repository directory
python src/spine/bin/run.py --config config/train_uresnet.yaml --source /path/to/data.h5Basic example:
# Necessary imports
from spine.config import load_config_file
from spine.driver import Driver
# Load configuration file
cfg_path = 'config/train_uresnet.yaml' # or your config file
cfg = load_config_file(cfg_path)
# Initialize driver class
driver = Driver(cfg)
# Execute model following the configuration regimen
driver.run()- Documentation is available at https://spine.readthedocs.io/latest/.
- Tutorials and examples can be found in the documentation.
Example configurations are available in the config folder:
| Configuration name | Model |
|---|---|
train_uresnet.yaml |
UResNet alone |
train_uresnet_ppn.yaml |
UResNet + PPN |
train_graph_spice.yaml |
GraphSpice |
train_grappa_shower.yaml |
GrapPA for shower fragments clustering |
train_grappa_track.yaml |
GrapPA for track fragments clustering |
train_grappa_inter.yaml |
GrapPA for interaction clustering |
To switch from training to inference mode, set trainval.train: False in your configuration file.
Key configuration parameters you may want to modify:
batch_size- batch size for training/inferenceweight_prefix- directory to save model checkpointslog_dir- directory to save training logsiterations- number of training iterationsmodel_path- path to checkpoint to load (optional)train- boolean flag for training vs inference modegpus- GPU IDs to use (leave empty '' for CPU)
For more information on storing analysis outputs and running custom analysis scripts, see the documentation on outputs (formatters) and analysis (scripts) configurations.
Basic usage with the spine command:
# Run training/inference directly
spine --config config/train_uresnet.yaml --source /path/to/data.h5
# Or run in background with logging
nohup spine --config config/train_uresnet.yaml --source /path/to/data.h5 > log_uresnet.txt 2>&1 &You can load a configuration file into a Python dictionary using:
from spine.config import load_config_file
# Load configuration file with SPINE's config loader
cfg = load_config_file('config/train_uresnet.yaml')A quick example of how to read a training log, and plot something
import pandas as pd
import matplotlib.pyplot as plt
fname = 'path/to/log.csv'
df = pd.read_csv(fname)
# plot moving average of accuracy over 10 iterations
df.accuracy.rolling(10, min_periods=1).mean().plot()
plt.ylabel("accuracy")
plt.xlabel("iteration")
plt.title("moving average of accuracy")
plt.show()
# list all column names
print(df.columns.values)Documentation for analysis tools and output formatting is available in the main documentation at https://spine.readthedocs.io/latest/.
bincontains utility scripts for data processingconfighas example configuration filesdocscontains documentation source filessrc/spinecontains the main package codetestcontains unit tests using pytest
Please consult the documentation for detailed information about each component.
The SPINE package includes comprehensive unit tests using pytest:
# Run all tests
pytest
# Run tests for a specific module
pytest test/test_data/
# Run with verbose output
pytest -vTest coverage tracking helps ensure code quality and identify untested areas. Coverage reports are automatically generated in our CI pipeline and uploaded to Codecov.
To check coverage locally:
# Run the coverage script (generates terminal, HTML, and XML reports)
./bin/coverage.sh
# Or run pytest with coverage flags directly
pytest --cov=spine --cov-report=term --cov-report=html
# View the HTML report
open htmlcov/index.htmlThe coverage configuration is defined in pyproject.toml under [tool.coverage.run] and [tool.coverage.report].
Before you start contributing to the code, please see the contribution guidelines.
The SPINE framework is designed to be extensible. To add a new model:
-
Data Loading: Parsers exist for various sparse tensor and particle outputs in
spine.io.core.parse. If you need fundamentally different data formats, you may need to add new parsers or collation functions. -
Model Implementation: Add your model to the
spine.modelpackage. Include your model in the factory dictionary inspine.model.factoriesso it can be found by the configuration system. -
Configuration: Create a configuration file in the
config/folder that specifies your model architecture and training parameters.
Once these steps are complete, you should be able to train your model using the standard SPINE workflow.